Identifying Circulating Tumor Cells by Image Analysis
Organization:
Funded by: | EU IMI supported CANCER-ID project |
PhD: | Leonie Zeune |
Supervisor: | Christoph Brune, Guus van Dalum (MCBP), Stephan van Gils, Leon Terstappen (MCBP) |
Collaboration: | Medical Cell Biophysics (MCBP), UTwente |
Description:
The EU IMI CANCER-ID (www.cancer-id.eu) project aims to validate blood-based biomarkers for cancer to determine the absence or presence of drug targets and to assess response to treatment. Within this project, we aim to develop an automatic detection and classification algorithm for Circulating Tumor Cells (CTCs) in blood probes using mathematical image analysis.
Applied Project background
Before CTCs can be established as a routine liquid biopsy in the clinic to guide therapy of cancer patients they will need to be present and isolated in sufficient numbers and their content readily accessible to determine whether certain therapies can be effective or not. An important consideration hereby is the lack of a unified approach to designate a cell as a CTC, in particular when different technology platforms are used.
Development of an automatic detection and classification algorithm for CTCs
The strong need for a unified, robust and automatic CTC definition will be addressed by the development of an Open Source CTC scoring algorithm. Microscopy images from different platforms can then be analyzed using the algorithm and, as a result, CTC candidates are presented to a reviewer. The main components of the algorithm are image normalization, segmentation and cell classification.
Mathematical imaging and inverse problems
To address those problems, our research focuses on variational methods in the field of mathematical imaging and inverse problems with biomedical applications. The mathematical research of Leonie Zeune is focused on multi-scale approaches for segmentation, Bregman distances, and nonlinear spectral methods. Besides, she is extending her work to classification and clustering methods. With regard to efficient and user-friendly implementations she is also working on primal-dual splitting techniques and methods for automatic parameter choices.
Publications:
Pictures:
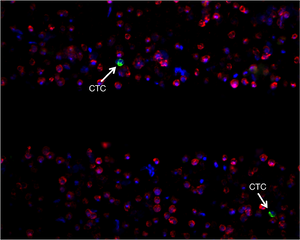
Figure 1: | |
| The blood of cancer patients is filtered and imaged with a fluorescence microscope. Cells are labeled using a white blood cell exclusion marker (red), an inclusion marker (green), and a DNA marker (blue).
|