Introduction
Cancer is the second leading cause of death. Globally, it is estimated that cancer is responsible for 9.6 million deaths in 2018 [1]. For almost half of the cancer patients, the disease is diagnosed at an advanced stage, which can greatly reduce the chances of survival [2]. Identification of early stage disease-specific alterations (biomarkers) could allow for enhanced detection of cancer. In the last decade, cancer biomarker research has been focusing on biomarkers representing genetic and epigenetic (e.a. phenotype changes that do not involve alterations in the DNA sequence) alterations associated with the development of cancer [3]. One of these epigenetic events includes DNA (hyper)methylation. DNA methylation is an epigenetic process involving the addition of a methyl group to cytosine. Hypermethylation in promoter regions can result in silencing of tumor suppressor genes, thereby driving carcinogenesis [4].
Background
Within the Weijerhorst project, which is a collaboration of several research groups within the University of Twente and researchers of AmsterdamUMC, the goal is set to sense cancer specific hmDNA in urine. The detection of hmDNA in urine is advantageous compared to conventional screening methods. Collection is painless, urine is produced in large amounts per patient per day, and no trained health-care professional is required for sampling. To determine DNA methylation, tools to distinguish between hmDNA and non-methylated DNA are needed. Raman spectroscopy has previously shown to be able to distinguish between methylated and nonmethylated cytosine [5-7]. This MSc assignment will focus on the development of an assay to distinguish hmDNA from normal genomic DNA via (surface enhanced) Raman/infra-red spectroscopy.
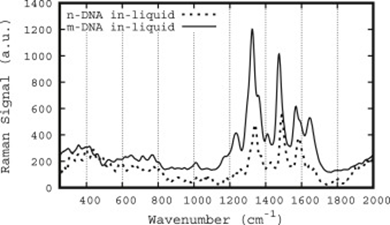
Figure 1: Comparison of methylated (excitation 258.2 nm) versus normal (excitation 268 nm) DNA Raman spectra. Taken From [8]
Research questions
This MSc assignment aims to address the following research questions:
- Which substrate is best to distinguish hmDNA from “normal” DNA
- Can gold nanoparticles be used as field enhancement tools for individual DNA molecule assessment?
- Can gold nanoparticle arrays be used to distinguish single hmDNA strands from “normal” DNA strands?
- Is it needed/possible/preferred to perform the spectroscopy on a (microfluidic) chip?
Contact person
References
[1] IARC, “Latest Global Cancer Data, 2018,” World Heal. Organ., no. September, pp. 13–15, 2018.
[2] M. H. Al-Azri, “Delay in cancer diagnosis: Causes and possible solutions,” Oman Medical Journal, vol. 31, no. 5, pp. 325–326, 2016.
[3] K. A. Calzone, “Genetic Biomarkers of Cancer Risk,” Semin. Oncol. Nurs., vol. 28, no. 2, pp. 122–128, May 2012.
[4] S. A. Qureshi, M. U. Bashir, and A. Yaqinuddin, “Utility of DNA methylation markers for diagnosing cancer,” International Journal of Surgery, vol. 8, no. 3. pp. 194–198, 2010.
[5] J. G. Kelly, G. M. Najand, and F. L. Martin, “Characterisation of DNA methylation status using spectroscopy (mid-IR versus Raman) with multivariate analysis,” J. Biophotonics, vol. 4, no. 5, pp. 345–354, May 2011.
[6] J. Kim, H. J. Park, J. H. Kim, B. Chang, and H.-K. Park, “Label-free Detection for a DNA Methylation Assay Using Raman Spectroscopy,” Chin. Med. J. (Engl)., vol. 130, no. 16, pp. 1961–1967, Aug. 2017.
[7] R. Daum, E. M. Brauchle, D. A. C. Berrio, T. P. Jurkowski, and K. Schenke-Layland, “Non-invasive detection of DNA methylation states in carcinoma and pluripotent stem cells using Raman microspectroscopy and imaging,” Sci. Rep., vol. 9, no. 1, p. 7014, Dec. 2019.
[8] Li, Ling, Shuang Fang Lim, Alexander Puretzky, Robert Riehn, and Hans D. Hallen. "DNA methylation detection using resonance and nanobowtie-antenna-enhanced Raman spectroscopy." Biophysical Journal 114, no. 11 (2018): 2498-2506.
